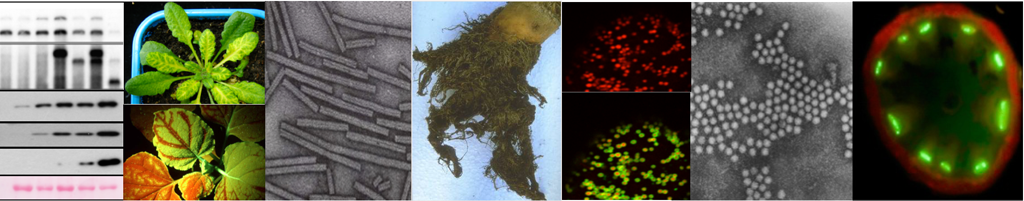
Early during viral infections, specific and non-specific interactions take place between both viral and host factors, leading to either progression or arrest of the infection. In particular mechanisms of plant defence can be countered by viruses encoding various pathogenicity factors. These viral factors can act by interfering with the innate defence pathways like RNA silencing or by deregulating the hormone signalling pathways to restrain or block the infection process. Our studies focus on the molecular mechanisms underlying pathogenicity, suppression of RNA silencing and viral movement as these viral functions appear to be closely coordinated during the multiplication cycle. The viruses studied belong to the Benyvirus and Polerovirus genera whose strategies of expression, spreading in the plant and vector-borne transmission are distinct.
Polerovirus movement in the phloem
Project manager:Véronique ZIEGLER-GRAFF
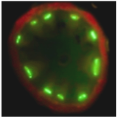
Poleroviruses are characterized by their obligate transmission by aphids and their phloem tropism. Recently we discovered a new protein P3a encoded by a cryptic gene conserved among poleroviruses. This small protein expressed from a non-conventional initiation codon is essential for viral long distance movement in the plant. Our objective is to decipher the mode of action of this small protein, in particular its interactions with the other viral proteins involved in viral movement.